Using AI to control energy for indoor agriculture
30 September 2024
Published online 19 February 2021
An online tool for exploring all the SARS-CoV-2 genomes sequenced so far will help researchers stay ahead of new variants.
Enlarge image
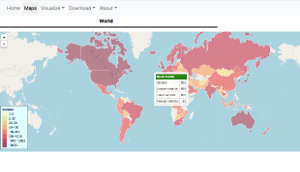
This is where computational scientists come to the fore. A team led by Intikhab Alam at King Abdullah University of Science and Technology (KAUST) in Saudi Arabia has developed an online system called CovMT that continually gathers data on newly sequenced genomes. Their website allows users to generate maps and timelines showing the evolution of the virus at global, continental or country-specific levels.
In May 2020, Alam and co-workers began intensive work on a computational pipeline to group individual SARS-CoV-2 genome samples into clades, lineages and more specific sets of mutations that they call mutation fingerprints.
“The main challenge initially was the lack of available genome sequencing data,” says Alam. However, the number of genomes increased exponentially in late 2020 when the World Health Organisation urged countries to increase sequencing efforts following the spread of mutations such as the highly transmissible UK variant, B.1.1.7. The new data are collected by GISAID (Global Initiative on Sharing Avian Influenza Data), an open hub for clinical, genetic and geographical data on viruses that was launched in 2008 in response to H5N1 bird flu.
“Thanks to GISAID, we are able to automatically download all new genomes daily. Once they are processed, the CovMT website is updated with the newest results and visualizations,” says Alam.
CovMT was able to confirm that the B.1.1.7 variant had acquired the E484K mutation, which helps the virus slip past the body’s immune defences and might compromise vaccination efforts. They are closely monitoring other concerning mutations, including L452R which is spreading in the US, and N501T in Oman, United Arab Emirates, Egypt and Jordan.
“I think this tracker is most helpful for tracking variants that might make vaccines less effective,” says Thomas Christie Williams, a researcher in evolutionary genetics at the University of Edinburgh who is involved in COVID-19 Genomics UK (COG-UK), a consortium that has provided nearly half of the world’s sequencing effort. “If these variants become widespread, approaches aiming to eradicate transmission of the virus – rather than merely preventing severe disease in the vulnerable – become much more difficult to achieve.”
Alam points out that Arab countries have some catching up to do in terms of sequencing efforts, although recent efforts have enabled CovMT to trace the emergence of the UK and South African variants in the UAE. “More significant and timely sequencing efforts are needed to detect these or other emerging variants in the Arab region,” he says. “Perhaps a joint consortium in the Gulf Cooperation Council, similar to COG-UK, may help.”
doi:10.1038/nmiddleeast.2021.16
Alam, I. et al. CovMT: an interactive SARS-CoV-2 mutation tracker, with a focus on critical variants. Lancet Infect Dis https://doi.org/10.1016/S1473-3099(21)00078-5 (2021).
Stay connected: